Resources
We believe in open science and in sharing meaningful research outputs beyond traditional publications. Our lab is committed to making protocols, data analysis and datasets open and easily accessible for anyone to use, reproduce and build upon.
2025

pCas9-Duo gRNA verification
[no publisher info]
·
11 Jan 2025
·
[no id info]
to screen for pCas9-Duo colonies positive for both guides.
2024

Design strategy for DiCre-mediated inducible gene knockout in the malaria parasite
protocols.io
·
24 Dec 2024
·
doi:10.17504/protocols.io.eq2ly68begx9/v1
This protocol describes a detailed strategy for inducibly knocking out a target gene in Plasmodium species using the Dimerisable Cre (DiCre) recombinase system, with a primary focus on Plasmodium falciparum.

Single-step assembly of double guide plasmid (pCas9-Duo) for gene-editing in Plasmodium
protocols.io
·
11 Jun 2024
·
doi:10.17504/protocols.io.n2bvjn9d5gk5/v1
This protocol describes a one-pot GoldenGate assembly of a new dual-guide targeting plasmid (pCas9-Duo) expressing two distinct guide RNAs in order to enhance the chances of a successful Cas9-mediated modification at a target region in Plasmodium falciparum.
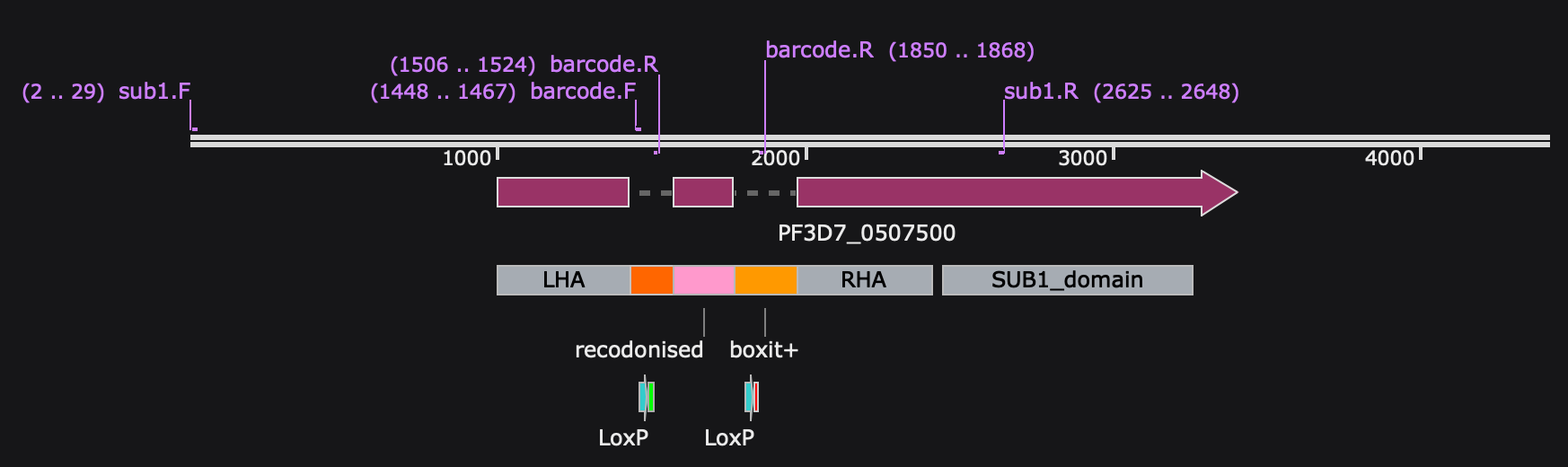
Plasmid and gene loci maps related to SHIFTiKO
Github
·
21 Jan 2024
·
[no id info]
2023
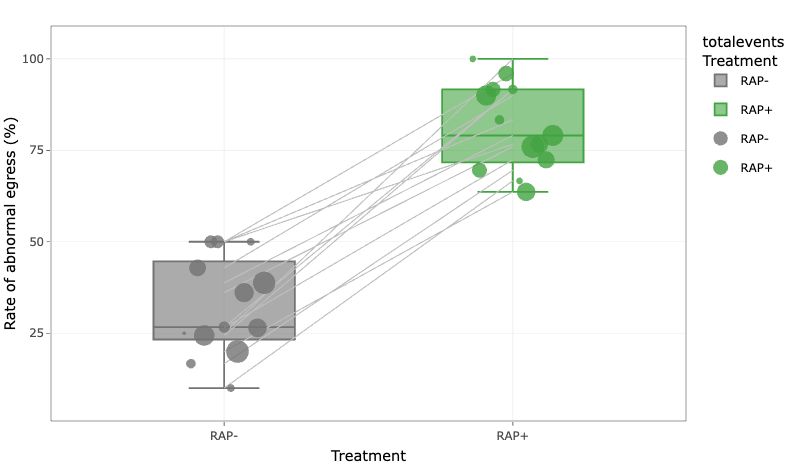
R notebook associated with the LCAT publication
[no publisher info]
·
29 May 2023
·
[no id info]
useful for exploring data on LCAT (PF3D7_0629300)-null mutants.
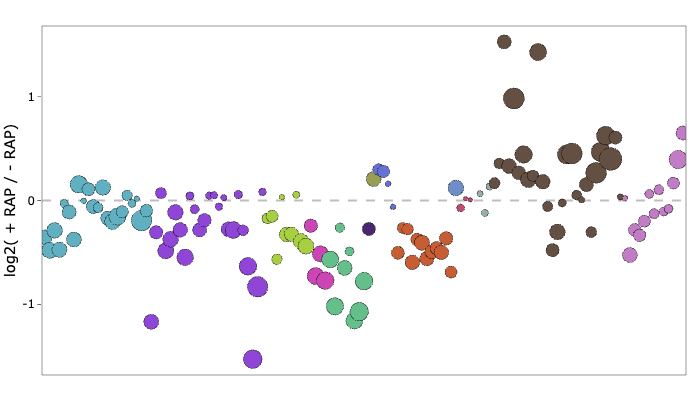
R notebook associated with the GDPD publication
[no publisher info]
·
14 May 2023
·
[no id info]
useful for exploring lipidomics data of GDPD (PF3D7_1406300)-null mutants.